UNIPPIN
UNIPPIN: An integrated human protein-protein interaction network
The group has been collecting and integrating protein interaction data from different major protein-protein interaction (PPI) databases and high-quality experimentally determined PPI data. By integrating all these data sources we compiled a Unified PPI network (UNIPPIN) for our analyses, ranging from extraction of topological features to mapping of genetic variants, protein-protein complex structures and protein-ligand interactions (drugs and allosteric ligands).
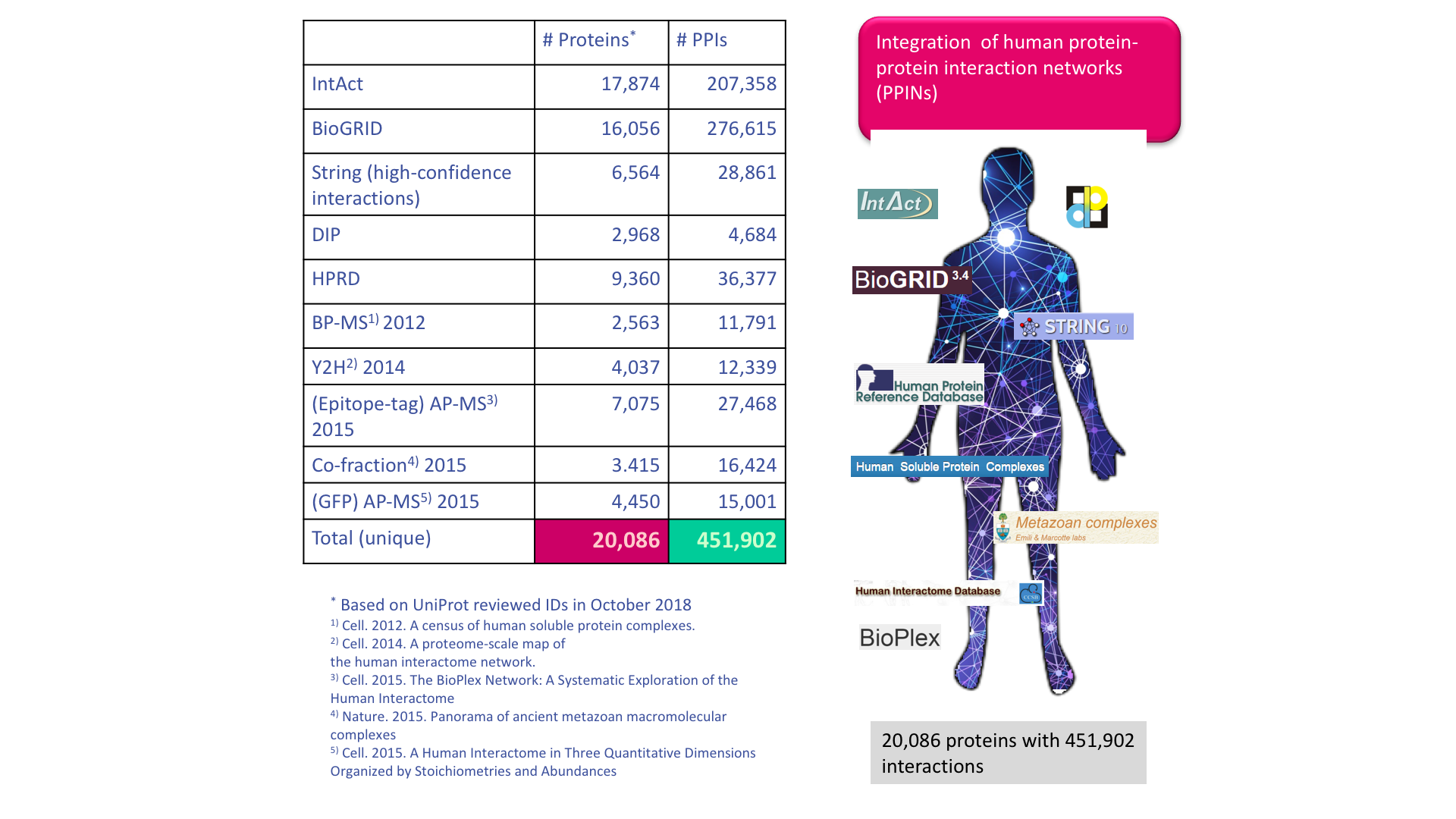
Representative publications on Protein-Protein interaction networks from the group:
- Anna Laddach, Sun Sook Chung and Franca Fraternali (2019) Prediction of Protein-Protein Interactions: Looking Through the Kaleidoscope. In: Ranganathan, S., Gribskov, M., Nakai, K. and Schönbach, C. (eds.), Encyclopedia of Bioinformatics and Computational Biology, vol. 2, pp. 834–848. Oxford: Elsevier.
- Laddach A, Ng JC, Chung SS, Fraternali F. (2018). Genetic variants and protein-protein interactions: a multidimensional network-centric view. Curr Opin Struct Biol. 2018 Jan 4;50:82-90. doi: 10.1016/j.sbi.2017.12.006. [Epub ahead of print]
- Chung SS, Laddach A, Thomas NSB, Fraternali F. (2018). Short loop motif profiling of protein interaction networks in acute myeloid leukaemia. bioRxiv. doi: 10.1101/306886
- Chung SS, Pandini A, Annibale A, Coolen AC, Thomas NS, Fraternali F. Bridging topological and functional information in protein interaction networks by short loops profiling. Sci Rep. 2015 Feb 23;5:8540. doi: 10.1038/srep08540.
- Carlin LM, Evans R, Milewicz H, Fernandes L, …, Fraternali F, Ameer-Beg S, Parker PJ, Thomas NS, Ng T. (2011). A targeted siRNA screen identifies regulators of Cdc42 activity at the natural killer cell immunological synapse. Sci Signal 4(201):ra81. doi: 10.1126/scisignal.2001729.
- Fernandes LP, Annibale A, Kleinjung J, Coolen AC, Fraternali F (2010). Protein networks reveal detection bias and species consistency when analysed by information-theoretic methods. PLoS One 5(8):e12083.